A Visual Introduction to Gap Statistics
A data expert shows us how to improve the findings of K-Means clustering in Python by employing Gap Statistics. Read on to get started!
Join the DZone community and get the full member experience.
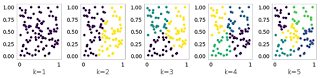
Join For FreeWe have previously seen how to implement K-Means. However, the results of this algorithm strongly rely on the choice of the parameter K. In this post, we will see how to use Gap Statistics to pick K in an optimal way. The main idea of the methodology is to compare the clusters inertia on the data to cluster and a reference dataset. The optimal choice of K is given by k for which the gap between the two results is maximum. To illustrate this idea, let’s pick as reference dataset a uniformly distributed set of points and see the result of K-Means increasing K:
import numpy as np
import matplotlib.pyplot as plt
from sklearn.datasets import make_blobs
from sklearn.metrics import pairwise_distances
from sklearn.cluster import KMeans
reference = np.random.rand(100, 2)
plt.figure(figsize=(12, 3))
for k in range(1,6):
kmeans = KMeans(n_clusters=k)
a = kmeans.fit_predict(reference)
plt.subplot(1,5,k)
plt.scatter(reference[:, 0], reference[:, 1], c=a)
plt.xlabel('k='+str(k))
plt.tight_layout()
plt.show()Let’s now do the same on a target dataset with three natural clusters:
plt.figure(figsize=(12, 3))
for k in range(1,6):
kmeans = KMeans(n_clusters=k)
a = kmeans.fit_predict(X)
plt.subplot(1,5,k)
plt.scatter(X[:, 0], X[:, 1], c=a)
plt.xlabel('k='+str(k))
plt.tight_layout()
plt.show()If we plot the inertia in both cases we note that on the reference dataset the inertia goes down very slowly while on the target dataset it assumes the shape of an elbow:
def compute_inertia(a, X):
W = [np.mean(pairwise_distances(X[a == c, :])) for c in np.unique(a)]
return np.mean(W)
def compute_gap(clustering, k_max=10, n_references=5):
reference_inertia = []
for k in range(1, k_max+1):
local_inertia = []
for _ in range(n_references):
clustering.n_clusters = k
assignments = clustering.fit_predict(reference)
local_inertia.append(compute_inertia(assignments, reference))
reference_inertia.append(np.mean(local_inertia))
ondata_inertia = []
for k in range(1, k_max+1):
clustering.n_clusters = k
assignments = clustering.fit_predict(X)
ondata_inertia.append(compute_inertia(assignments, X))
gap = np.log(reference_inertia)-np.log(ondata_inertia)
return gap, np.log(reference_inertia), np.log(ondata_inertia)
gap, reference_inertia, ondata_inertia = compute_gap(KMeans())
plt.plot(range(1, k_max+1), reference_inertia,
'-o', label='reference')
plt.plot(range(1, k_max+1), ondata_inertia,
'-o', label='data')
plt.xlabel('k')
plt.ylabel('log(inertia)')
plt.show()We can now compute the Gap Statistics for each K computing the difference of the two curves shown above:
plt.plot(range(1, k_max+1), gap, '-o')
plt.ylabel('gap')
plt.xlabel('k')It’s easy to see that the Gap is maximum for K=3, just the right choice for our target dataset.
Published at DZone with permission of Giuseppe Vettigli, DZone MVB. See the original article here.
Opinions expressed by DZone contributors are their own.




Comments