In-Memory Technologies: Meeting Healthcare's Fast Data Challenges (Part 2)
Learn about a healthcare case study from a company called e-Therapeutics, which specializes in drug discovery and development, and see how they used Apache Ignite.
Join the DZone community and get the full member experience.
Join For FreeThis is the second in a two-part series on the use of fast data in healthcare and how in-memory technologies such as Apache Ignite can meet the requirements and challenges of the healthcare industry. Part 1 focused on identifying some of the key challenges in Healthcare. In Part 2, we will discuss a healthcare case study and learn how Apache Ignite and GridGain solved a customer's problem.
Introduction
The customer case study we will discuss is from a company called e-Therapeutics. The company, founded in the United Kingdom in 2003, specializes in drug discovery and development. In particular, it is focused on finding treatments for diseases such as cancer and diseases that cause degeneration of the nervous system such as Parkinson's and Alzheimer's.
The Business Challenges
The first challenge for e-Therapeutics was in the area of network pharmacology. This is where a specific network of proteins associated with a particular disease is analyzed and identified. The next step is to identify multiple intervention points to disrupt the network of proteins. The goal is to discover drug molecules that would provide the best disruption of a protein network. Therefore, significant data may need to be stored and analyzed. In the previous article, we discussed Apache Ignite's architecture and its ability to scale.
The second challenge was in the area of computational analysis of the disease cells. To save time and resources, multiple analyses need to be performed. These analyses involve a varying range of parameters. The analyses are also very compute-intensive. Therefore, parallelism offered a good solution. Apache Ignite's In-Memory Compute Grid allows the execution of distributed computations in parallel to obtain high performance, low latency, and linear scalability. Ignite's Compute Grid provides a rich set of APIs that allow users to distribute computations and data processing across multiple computers in a cluster. Collocating data and bringing the processing to the server where the data reside also provides benefits, such as reduced network traffic.
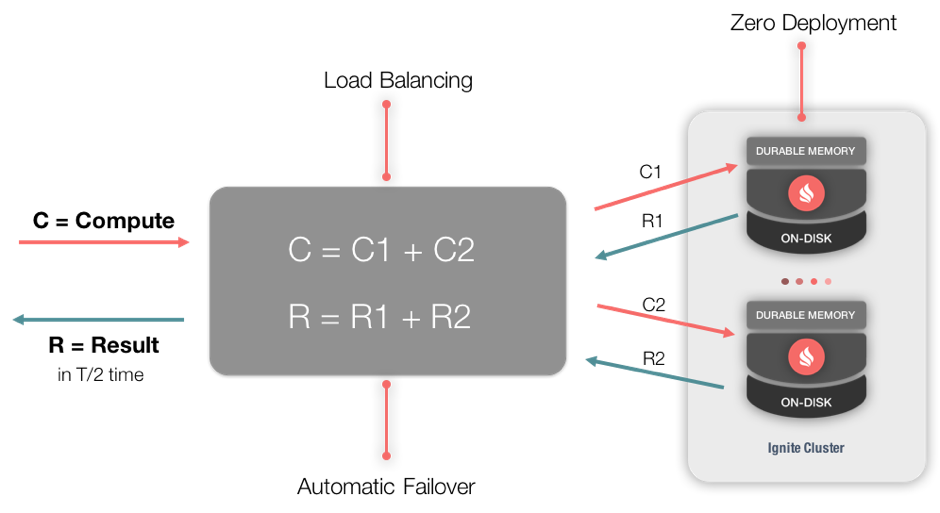
Figure 1: Compute Grid
Figure 1 shows an example Apache Ignite Compute Grid with two servers. Task C is split into multiple jobs (C1, C2). These jobs are sent to the two servers, respectively. The results received (R1, R2) from the two servers are combined (R) and returned to the client.
The Benefits of using Apache Ignite
e-Therapeutics opted to use GridGain's solution for its Network Pharmacology platform. GridGain's technology is built upon Apache Ignite and provides resources, support, and enterprise capabilities.
The first benefit for e-Therapeutics was improved performance. Originally, the company started with a cluster consisting of 20 nodes on a 20-core server. This later grew to 100 nodes on five servers (Figure 2).
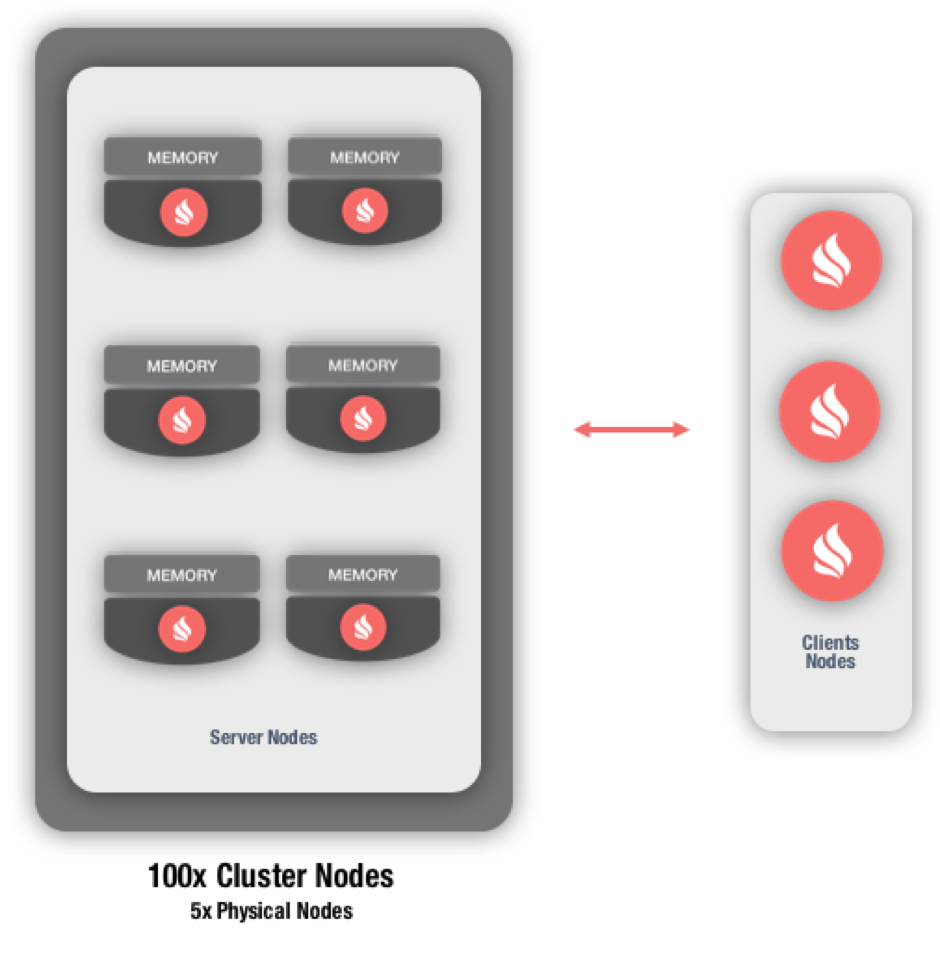
Figure 2: e-Therapeutics platform
By using parallelism, the company saw a speed increase of nearly two orders of magnitude when compared to the old non-parallelized version. Improved performance meant that analyses could be completed in hours and minutes, while also allowing new projects to be undertaken that were previously infeasible.
The second benefit for e-Therapeutics was the improved productivity of staff members. Disease biology specialists and researchers are not computational informatics specialists. So, a web-based interface connecting to microservices was developed to allow access to the new platform. Staff members could run analyses without having to work from a command line. Furthermore, there was no need to consult with a computational informatics specialist. Scientists could also now work on multiple projects and achieve far more in less time. For example, over a period of 18 months, e-Therapeutics was able to run ten concurrent discovery projects with a small team of scientists. As a result of the faster processing and improved productivity, drug discoveries could be moved into testing phases much faster.
The third benefit for e-Therapeutics was peace of mind. Apache Ignite is a top-level project at the Apache Software Foundation (ASF). ASF has a great reputation for providing stability and longevity. Many projects have been hosted by ASF over a long period of time, providing high-quality, community-driven software.
Summary
e-Therapeutics provides a specialized approach to network biology using a computer-based drug discovery platform built upon Apache Ignite. The original problems for the company were the time required for computational analyses to be performed and the inability of existing algorithms to be parallelized. The solutions were to develop a new platform based on Apache Ignite that used parallelism and provided nearly two-orders of magnitude performance improvement, enabling work to be completed in hours and minutes rather than weeks and days.
Published at DZone with permission of Akmal Chaudhri. See the original article here.
Opinions expressed by DZone contributors are their own.

Comments