Doing Residual Analysis Post Regression in R
In this post, we take a deep dive into the R language by exploring residual analysis and visualizing the results with R.
Join the DZone community and get the full member experience.
Join For FreeResiduals are essentially gaps that are left when a given model, in this case, linear regression, does not fit the given observations completely.
A close analogy for residual analysis is found in medical pathology. What remains post-metabolism usually becomes an indicator of what was processed and absorbed versus what was not so.
Residuals are the difference between the original value to be modeled and the estimator of original value that came as a result of your model.
Residual error = Y- Y-hat, where Y is the original value and Y-hat is the computed value.
I would expect this error to be as close to zero as possible and will be interested in finding any outliers or aberrations thereof.
A quick way to find outliers is to use either standardized or studentized residuals. The first is simply scaling the residuals relative to their standard deviation and, hence, anything that lies beyond 3 standard deviations can be considered to be an aberration.
## standardize residuals - scale residuals relative to their standard deviation
residueStandard<-rstandard(lmfit)
df[residueStandard>3,]Here is what you get:
days.instant days.atemp days.hum days.windspeed days.casual
442 442 0.505046 0.755833 0.110704 3155
456 456 0.421708 0.738333 0.250617 2301
463 463 0.426129 0.254167 0.274871 3252
470 470 0.487996 0.502917 0.190917 2795
471 471 0.573875 0.507917 0.225129 2846
505 505 0.566908 0.456250 0.083975 3410
512 512 0.642696 0.732500 0.198992 2855
513 513 0.641425 0.697083 0.215171 3283
533 533 0.594708 0.504167 0.166667 2963
624 624 0.585867 0.501667 0.247521 3160
645 645 0.538521 0.664167 0.268025 3031
659 659 0.472842 0.572917 0.117537 2806Naturally, I would expect my model to be unbiased, at least in intention, and hence any leftovers on either side of the regression line that did not make it on the line are expected to be random, i.e. without any particular pattern.
That is, I expect my residual error distributions to follow a bland, normal distribution.
In R, you can do this elegantly with just two lines of code.
1. Plot a histogram of residuals
2. Add a Quantile-Quantile plot with a line that passes through, namely, the first and third quantiles.
hist(lmfit$residuals)
qqnorm(lmfit$residuals);qqline(lmfit$residuals)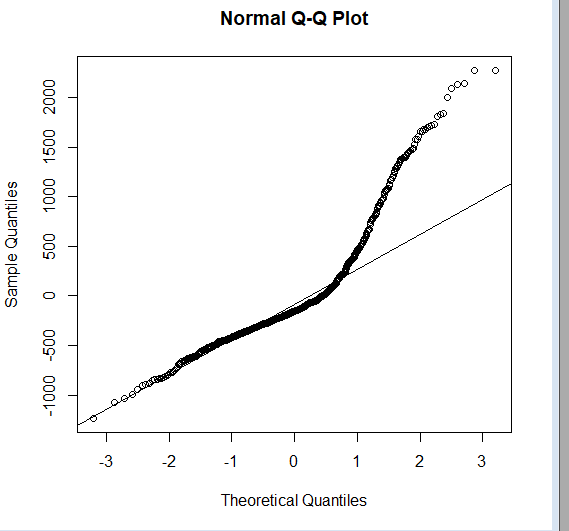
So we know that the plot deviates from normal (represented by the straight line).
But that binary aspect of information is seldom enough. So, we should check the skewness and kurtosis to know the dispersion of the distribution.
First, we will compute the skewness; we will use a simple Galton's skewness formula.
## check on symmetry of distribution: skewness
summary<-summary(lmfit$residuals)
Q1<-summary[[2]]
Q2<-summary[[3]]
Q3<-summary[[5]]
skewness<-((Q1-Q2)+(Q3-Q2))/(Q3-Q1)
skewness
[1] 0.2665384This quantifies which side the distribution has swayed to (the right side or left side, or if it's perfectly symmetrical or even).
Another measuring aspect is Kurtosis, which will tell you whether the distribution is assembled towards the center (+ve value) or away from it. (-ve value). Kurtosis is a metric that quantifies the possibility of outliers. A positive value indicates their existence.
kurtosis(lmfit$residuals)
[1] 2.454145Next, apart from visualizing the distribution of residual errors, I would also like to find whether the residuals are correlated with each other.
For a model to account for all the variation in observations, residual errors must occur at random and be uncorrelated with each other.
Durbin-Watson test allows you to test the independence of residuals from each other.
dwt(lm(df$days.windspeed~df$days.atemp+df$days.hum),method='resample', alternative='two.sided')
lag Autocorrelation D-W Statistic p-value
1 0.2616792 1.475681 0
Alternative hypothesis: rho != 0The statistic value needs to be compared with lower and upper limits established by the critical values table, for a given degree of freedom and number of observations. In this case, the range is [1.55,1.67].
Since the computed D-W statistic falls below the lower value of this range, we reject the null hypothesis that residuals are not correlated. Hence, an alternate hypothesis that there may be a correlation between residuals comes into play.
Visually, this can be learned by studying how a model falls short when attempting to capture increases in the value of the original Y value. As this happens, the residuals appear proportional to Y, as Y increases.
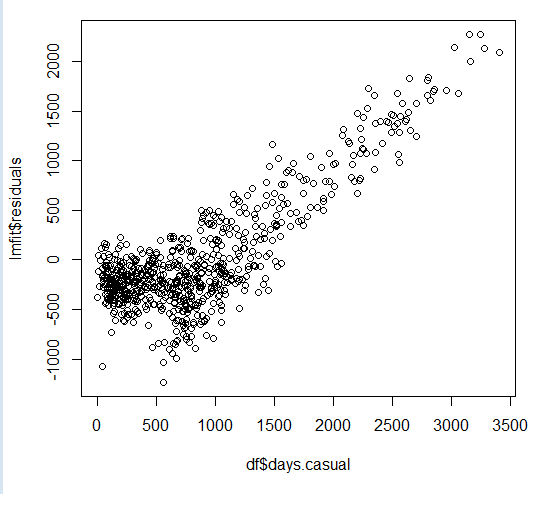
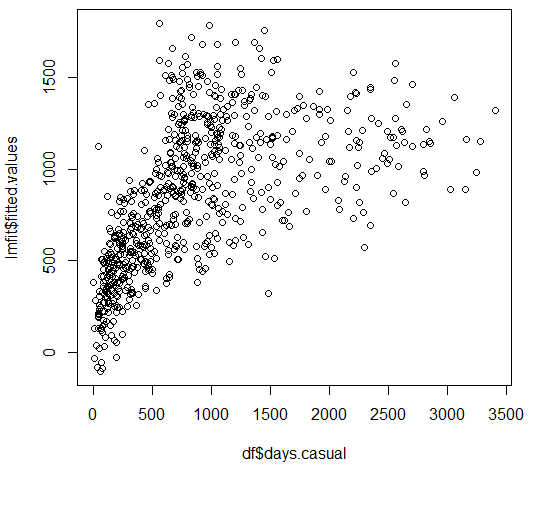
Compare this with plotting the fitted Y-hat value against Y. Where Y-hat values tend to fall behind, residuals appear to run along with Y and hence the past residual value appears to be a better predictor of present values as they appear to continue along a fixed slope value. And correlation shows up in Darwin-Watson tests that try to compare and contrast the sum square of the difference between residual errors and their previous value(s) in relation to the sum of a given residual error over all observations.
Correlation between residual errors is thus like failing on a cardio test. If your model simply can't keep up with the rapid changes in Y, it will start looking more and more like an exasperated runner, with the count of lost steps appearing to be more and more correlated than the actual steps taken by the runner.
Reconstructing the subconscious influence your linear regression model was under.
It is important to remember that when you run a linear regression on any given set of observations, each value of THE computed estimator (symbolically represented as Y-hat) of THE dependent variable (similarly represented as Y), is not only dependant on the current value (say, with the observation), but also dependant on each observation.
After all, your multiple linear regression models computed the coefficients for each contributing factor, only after minimizing the error between the actual value of dependent variable Y and the computed estimator of Y (Y-hat) on all observations.
Mathematically, this can be represented by the simple weighted average model as follows.
Yhat(j)= w1j*Y(1)+w2j*Y(2)+w3j*Y(3)+w4j*Y(4)+.....+wnj*Y(n);
where n is number of observations and j is one particular estimated value. If you want to think of this in matrix notation, matrix W will have dimensions [n,n] where n is the number of observations. Similarly, Y represented in matrix form will have dimensions [n,1], and hence Y-hat[j] is represented as a matrix because it is obtained as a product of two matrices (W and Y) and will have the dimensions [n,1], which are the same as Y.
So, each value of the computed estimator can be expressed as a weighted average of all original Y values, indicating to what extent a given computed estimator is influenced by each original Y value.
To put it more simply, if I see a bunch of observations scattered here and there, and I have to somehow draw a straight line that passes through them with as little bias as possible, that is, if I must leave out some observations, I must do so equally on either side of this regression line.
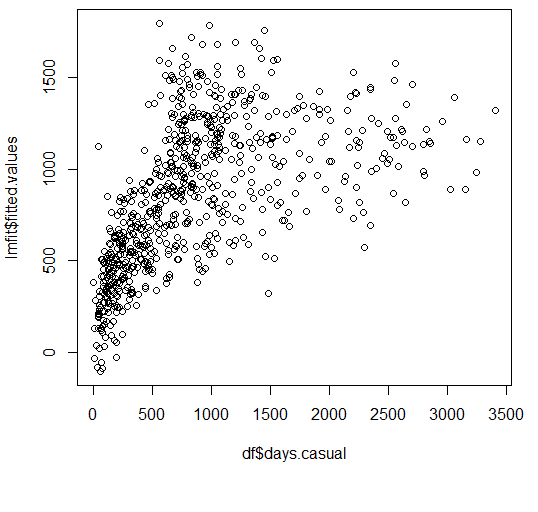
But while doing so, I (or the regression program of choice) subconsciously estimated to what extent the slope of my regression line was influenced by each observation, irrespective of where it belonged.
All I am doing now is back calculating such influences in retrospective. How much weight did each observation carry in order to sway my regression line to either include them or exclude them?
The statistical term for weights is hat values because they bridge computed estimators and their original counterparts.
This is how it is computed in R.
modelmatrix<-model.matrix(lmfit)
hatvalues<-hat(modelmatrix)First, we get the model in a matrix form. Then we compute the hat values.
Alternatively, you can use the following function to get a similar result.
hatvalues<-lm.influence(lmfit)$hatLet's think about the limits that can be imposed on each weight.
Obviously, the lowest possible value for the weight will be equal to the possibility when all original Y values contribute equally (since they must contribute something for the linear regression program to estimate the coefficients by working and optimizing through all observations).
In this case, the lower limit of its range will be 1/n, where n is the total number of observations. In any case, since n will always be +ve, the weights will always be <1.
Now try to sum up your hat values and you will see an interesting result...
sum(hatvalues)
[1] 4They equal the number of contributing factors your linear regression model has considered for computation +1.
For instance, in the example dataset, we have three contributing factors, namely: temperature, humidity, and wind speed.
lmfit$coefficients
(Intercept) df$days.atemp df$days.hum df$days.windspeed
490.2956 2312.0163 -875.6955 -992.1411Next, how do we find the most significant or influencing observations?
An elegant way will be to just:
segment hat values into quantiles .
apply the 95% criterion to filter the most aberrant values.
apply this filtering criterion back on observations.
R allows you to all of this in one step!
df[hatvalues>quantile(hatvalues,0.95),]We can get specific observations that satisfy this criterion.
days.instant days.atemp days.hum days.windspeed days.casual
9 9 0.1161750 0.434167 0.361950 54
21 21 0.1578330 0.457083 0.353242 75
22 22 0.0790696 0.400000 0.171970 93
26 26 0.2036000 0.862500 0.293850 34
32 32 0.2345300 0.829565 0.053213 47
36 36 0.2430580 0.929167 0.161079 100
45 45 0.3983500 0.375833 0.417908 208
50 50 0.3914040 0.187917 0.507463 532
65 65 0.3662520 0.948261 0.343287 114
69 69 0.3856680 0.000000 0.261877 46
90 90 0.2575750 0.918333 0.217646 179
94 94 0.5429290 0.426250 0.385571 734
95 95 0.3983500 0.642083 0.388067 167
106 106 0.4254920 0.888333 0.340808 121
153 153 0.6439420 0.305000 0.292287 736
239 239 0.6355560 0.850000 0.375617 226
249 249 0.5152000 0.886957 0.343943 204
293 293 0.4665250 0.636250 0.422275 471
302 302 0.2279130 0.882500 0.351371 57
341 341 0.4002460 0.970417 0.266175 50
368 368 0.1262750 0.441250 0.365671 89
383 383 0.2752540 0.443333 0.415429 109
388 388 0.2430580 0.911250 0.110708 145
408 408 0.1016580 0.464583 0.409212 73
421 421 0.2556750 0.395833 0.421642 317
433 433 0.5246040 0.567500 0.441563 486
434 434 0.3970830 0.407083 0.414800 447
463 463 0.4261290 0.254167 0.274871 3252
465 465 0.4766380 0.317500 0.358196 905
478 478 0.3895040 0.835417 0.344546 120
543 543 0.5947040 0.373333 0.347642 1077
627 627 0.5650670 0.872500 0.357587 371
667 667 0.4677710 0.694583 0.398008 998
668 668 0.4394000 0.880000 0.358200 2
694 694 0.2487420 0.404583 0.376871 532
722 722 0.2361130 0.441250 0.407346 205
726 726 0.2203330 0.823333 0.316546 9Finally, you can visualize them with color coding.
##visualize
plot(df$days.casual,lmfit$fitted.values,col=ifelse(hatvalues>quantile(hatvalues,0.95),'red','blue'))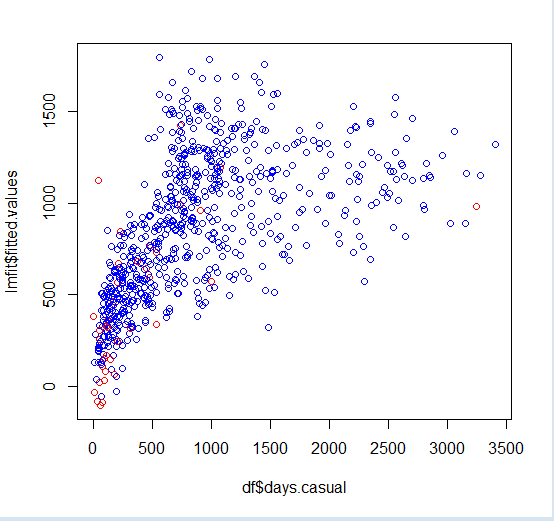
That's it! This is a simple overview of doing residual analysis and why it's important.
The link to the data set and model assumptions are mentioned in the references section.
Opinions expressed by DZone contributors are their own.

Comments